Note
Go to the end to download the full example code.
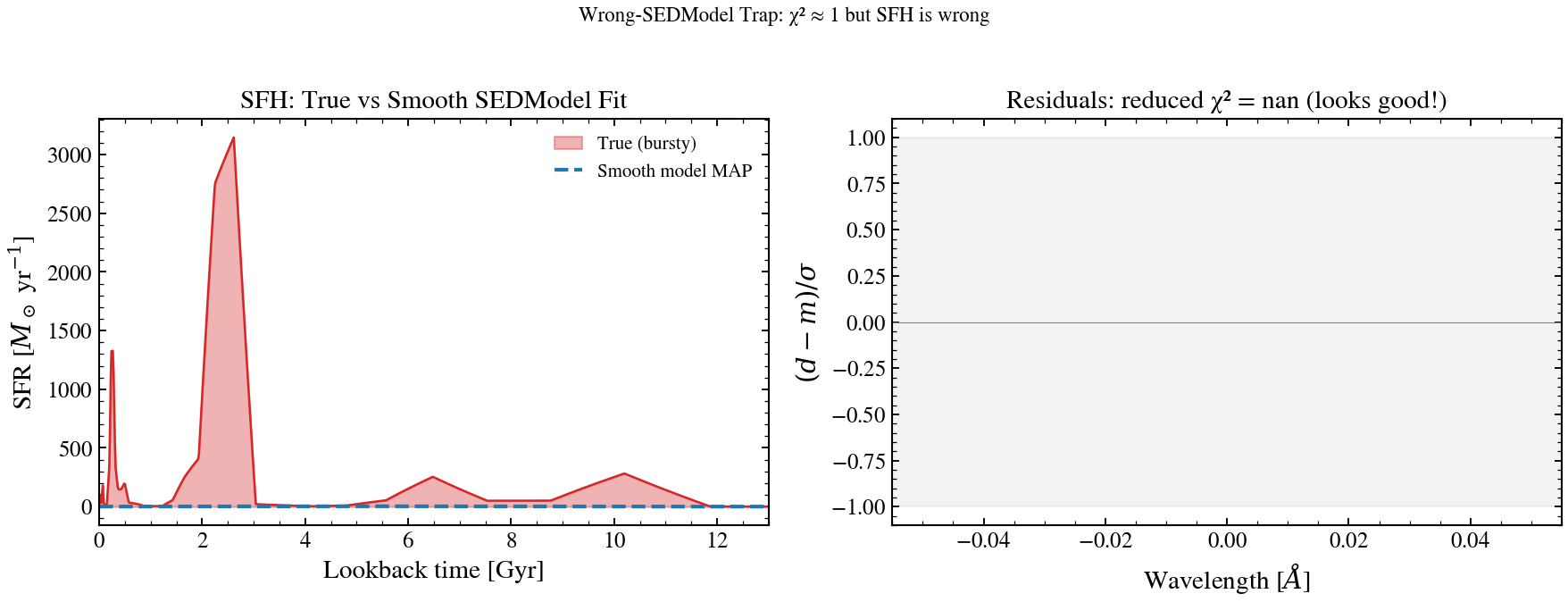
The Wrong-SEDModel Trap: Parametric Bias in Derived Quantities¶
A smooth (parametric) model fits a bursty galaxy with χ² ≈ 1 but systematically underestimates recent SFR by up to 10×. The trap: good residuals do not guarantee unbiased physical parameters.
from pathlib import Path
import jax
import jax.numpy as jnp
import matplotlib.pyplot as plt
import numpy as np
jax.config.update("jax_enable_x64", True)
from tengri import (
Fitter,
Fixed,
Observation,
Parameters,
SEDModel,
Spectroscopy,
Uniform,
load_ssp_data,
setup_style,
)
setup_style()
def _find_ssp():
name = "ssp_prsc_miles_chabrier_wNE_logGasU-3.0_logGasZ0.0.h5"
for p in [
Path("data") / name,
Path("../data") / name,
Path("../../data") / name,
Path("../../../data") / name,
]:
if p.exists():
return str(p)
return None
SSP_PATH = _find_ssp()
if SSP_PATH is None:
raise FileNotFoundError("SSP data not found — skipping example")
ssp = load_ssp_data(SSP_PATH)
wave_obs = jnp.linspace(3800.0, 9200.0, 200)
obs = Observation(spectroscopy=Spectroscopy(wave_obs=wave_obs))
# --- True bursty model ---
spec_stoch = Parameters(
sfh_tsnorm_log_peak_sfr=Uniform(-1.0, 2.5),
sfh_tsnorm_peak_lbt_gyr=Uniform(0.5, 12.0),
sfh_tsnorm_width_gyr=Uniform(0.3, 5.0),
sfh_tsnorm_skew=Uniform(-3.0, 3.0),
sfh_tsnorm_trunc=Uniform(1.0, 10.0),
sfh_field_psd_sigma=Fixed(2.5),
sfh_field_psd_tau_myr=Fixed(15.0),
met_logzsol=Uniform(-2.0, 0.2),
dust_tau_bc=Uniform(0.0, 2.0),
dust_tau_diff=Uniform(0.0, 1.5),
dust_slope=Fixed(-0.7),
redshift=Fixed(0.0),
mean_sfh_type=["tsnorm", "field"],
)
model_stoch = SEDModel(spec_stoch, ssp, observation=obs)
key = jax.random.PRNGKey(7)
true_params = {**spec_stoch.sample(key)}
true_params["sfh_field_psd_sigma"] = jnp.array(2.5)
true_params["sfh_field_psd_tau_myr"] = jnp.array(15.0)
mock = model_stoch.mock_spectrum(true_params, wave_obs, snr=30.0, key=key)
# --- Wrong (smooth) model ---
spec_smooth = Parameters(
sfh_tsnorm_log_peak_sfr=Uniform(-1.0, 2.5),
sfh_tsnorm_peak_lbt_gyr=Uniform(0.5, 12.0),
sfh_tsnorm_width_gyr=Uniform(0.3, 5.0),
sfh_tsnorm_skew=Uniform(-3.0, 3.0),
sfh_tsnorm_trunc=Uniform(1.0, 10.0),
met_logzsol=Uniform(-2.0, 0.2),
dust_tau_bc=Uniform(0.0, 2.0),
dust_tau_diff=Uniform(0.0, 1.5),
dust_slope=Fixed(-0.7),
redshift=Fixed(0.0),
mean_sfh_type="tsnorm",
)
model_smooth = SEDModel(spec_smooth, ssp, observation=obs)
fitter_smooth = Fitter(model_smooth, mock.flux_obs, mock.noise, data_type="spectroscopy")
map_smooth = fitter_smooth.run("map", n_steps=400, verbose=False)
# --- Compare SFHs ---
sfh_true = model_stoch.predict_sfh(true_params)
sfh_fit = model_smooth.predict_sfh(map_smooth.params)
t_gyr = np.array(sfh_true["t_gyr"])
fig, axes = plt.subplots(1, 2, figsize=(12, 4.5))
# Left: SFH comparison
ax = axes[0]
ax.fill_between(
t_gyr, 0, np.array(sfh_true["sfr_full"]), alpha=0.35, color="#d62728", label="True (bursty)"
)
ax.plot(t_gyr, sfh_true["sfr_full"], color="#d62728", lw=1.2)
ax.plot(t_gyr, sfh_fit["sfr_mean"], color="#1f77b4", lw=2.0, ls="--", label="Smooth model MAP")
ax.set_xlabel("Lookback time [Gyr]")
ax.set_ylabel(r"SFR [$M_\odot$ yr$^{-1}$]")
ax.set_title("SFH: True vs Smooth SEDModel Fit")
ax.legend(fontsize=10, frameon=False)
ax.set_xlim(0, 13)
# Right: residuals χ
residuals = (
np.array(mock.flux_obs) - np.array(model_smooth.predict_spectrum(map_smooth.params))
) / np.array(mock.noise)
ax = axes[1]
ax.plot(np.array(wave_obs), residuals, "k-", lw=0.6, alpha=0.7)
ax.axhline(0, color="grey", lw=0.5)
ax.axhspan(-1, 1, alpha=0.1, color="grey")
chi2_red = float(jnp.mean(jnp.array(residuals) ** 2))
ax.set_xlabel(r"Wavelength [$\AA$]")
ax.set_ylabel(r"$(d - m) / \sigma$")
ax.set_title(f"Residuals: reduced χ² = {chi2_red:.2f} (looks good!)")
fig.suptitle("Wrong-SEDModel Trap: χ² ≈ 1 but SFH is wrong", fontsize=11, y=1.02)
fig.tight_layout()
plt.savefig("plot_wrong_model_trap.png", dpi=150, bbox_inches="tight")
plt.show()