Note
Go to the end to download the full example code.
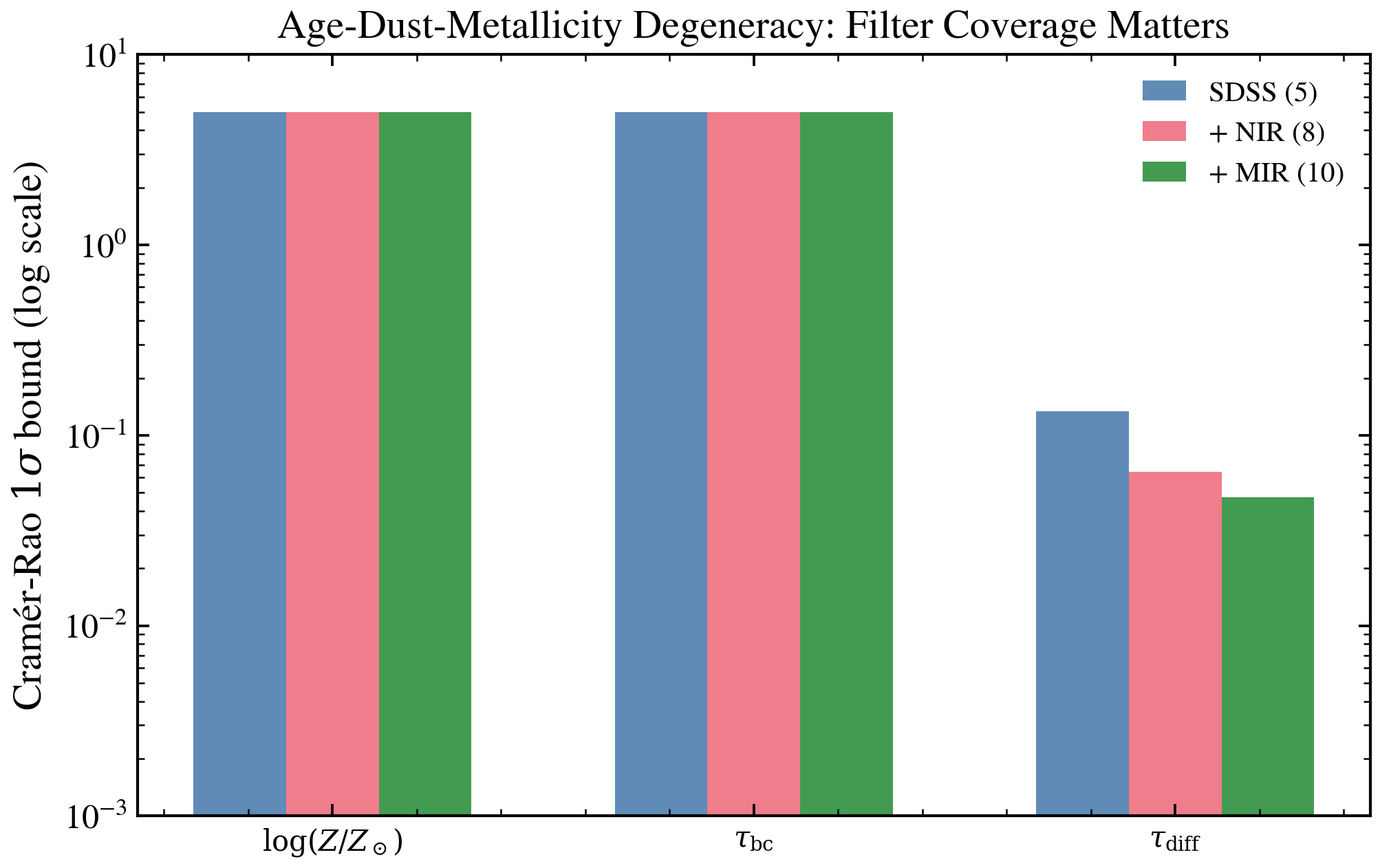
Age-Dust-Metallicity Degeneracy: Fisher Analysis¶
The Cramér-Rao bound from the Fisher Information Matrix shows that SDSS 5-band photometry alone cannot separately constrain age, dust, and metallicity. Adding NIR or MIR bands breaks the degeneracy by factors of 2–5×, quantifying the information gain from multiwavelength coverage.
from pathlib import Path
import jax
import jax.numpy as jnp
import matplotlib.pyplot as plt
import numpy as np
jax.config.update("jax_enable_x64", True)
from tengri import (
Fixed,
Observation,
Parameters,
Photometry,
SEDModel,
Uniform,
load_ssp_data,
setup_style,
)
from tengri.analysis.diagnostics.fisher import compute_fisher_matrix, fisher_parameter_errors
setup_style()
def _find_ssp():
name = "ssp_prsc_miles_chabrier_wNE_logGasU-3.0_logGasZ0.0.h5"
for p in [
Path("data") / name,
Path("../data") / name,
Path("../../data") / name,
Path("../../../data") / name,
]:
if p.exists():
return str(p)
return None
SSP_PATH = _find_ssp()
if SSP_PATH is None:
raise FileNotFoundError("SSP data not found — skipping example")
ssp = load_ssp_data(SSP_PATH)
_FILTER_DIR = next(
(
str(d)
for d in [
Path("data/filters"),
Path("../data/filters"),
Path("../../data/filters"),
Path("../../../data/filters"),
]
if d.exists()
),
"data/filters",
)
spec = Parameters(
sfh_tsnorm_log_peak_sfr=Uniform(-1.0, 2.5),
sfh_tsnorm_peak_lbt_gyr=Uniform(0.5, 12.0),
sfh_tsnorm_width_gyr=Uniform(0.3, 5.0),
sfh_tsnorm_skew=Uniform(-3.0, 3.0),
sfh_tsnorm_trunc=Uniform(1.0, 10.0),
met_logzsol=Uniform(-2.0, 0.2),
dust_tau_bc=Uniform(0.0, 2.0),
dust_tau_diff=Uniform(0.0, 1.5),
dust_slope=Fixed(-0.7),
redshift=Fixed(0.1),
mean_sfh_type="tsnorm",
)
key = jax.random.PRNGKey(42)
true_params = {
**spec.sample(key),
"met_logzsol": jnp.array(-0.3),
"dust_tau_bc": jnp.array(0.8),
"dust_tau_diff": jnp.array(0.4),
}
FILTER_SETS = {
"SDSS (5)": ["sdss_u", "sdss_g", "sdss_r", "sdss_i", "sdss_z"],
"+ NIR (8)": [
"sdss_u",
"sdss_g",
"sdss_r",
"sdss_i",
"sdss_z",
"2mass_j",
"2mass_h",
"2mass_ks",
],
"+ MIR (10)": [
"sdss_u",
"sdss_g",
"sdss_r",
"sdss_i",
"sdss_z",
"2mass_j",
"2mass_h",
"2mass_ks",
"wise_w1",
"wise_w2",
],
}
fisher_params = ["met_logzsol", "dust_tau_bc", "dust_tau_diff"]
PARAM_LABELS = [r"$\log(Z/Z_\odot)$", r"$\tau_{\rm bc}$", r"$\tau_{\rm diff}$"]
COLORS_BAR = ["#4477AA", "#EE6677", "#228833"]
sigmas = {}
for fname, filters in FILTER_SETS.items():
try:
obs = Observation(photometry=Photometry.from_names(filters, cache_dir=_FILTER_DIR))
mdl = SEDModel(spec, ssp, observation=obs)
phot = jnp.abs(mdl.predict_photometry(true_params))
noise = phot / 20.0
fim, _ = compute_fisher_matrix(
mdl, true_params, noise, data_type="photometry", param_names=fisher_params
)
errs = np.array(fisher_parameter_errors(fim))
# Unconstrained directions → clip to prior scale for visibility.
errs = np.where(np.isfinite(errs) & (errs > 0), errs, 5.0)
sigmas[fname] = np.minimum(errs, 5.0)
except Exception as e:
print(f"[{fname}] skipped: {e}")
if not sigmas:
raise RuntimeError("Fisher computation failed — check filter availability")
x = np.arange(len(fisher_params))
width = 0.22
fig, ax = plt.subplots(figsize=(7, 4.5))
for i, (fname, sigma_arr) in enumerate(sigmas.items()):
ax.bar(x + (i - 1) * width, sigma_arr, width, label=fname, color=COLORS_BAR[i], alpha=0.85)
ax.set_yscale("log")
ax.set_ylim(1e-3, 1e1)
ax.set_xticks(x)
ax.set_xticklabels(PARAM_LABELS, fontsize=10)
ax.set_ylabel(r"Cramér-Rao $1\sigma$ bound (log scale)")
ax.set_title("Age-Dust-Metallicity Degeneracy: Filter Coverage Matters")
ax.legend(fontsize=10, frameon=False)
fig.tight_layout()
plt.savefig("plot_fisher_degeneracy.png", dpi=150, bbox_inches="tight")
plt.show()