Note
Go to the end to download the full example code.
Population VI scaling: time, memory, and convergence¶
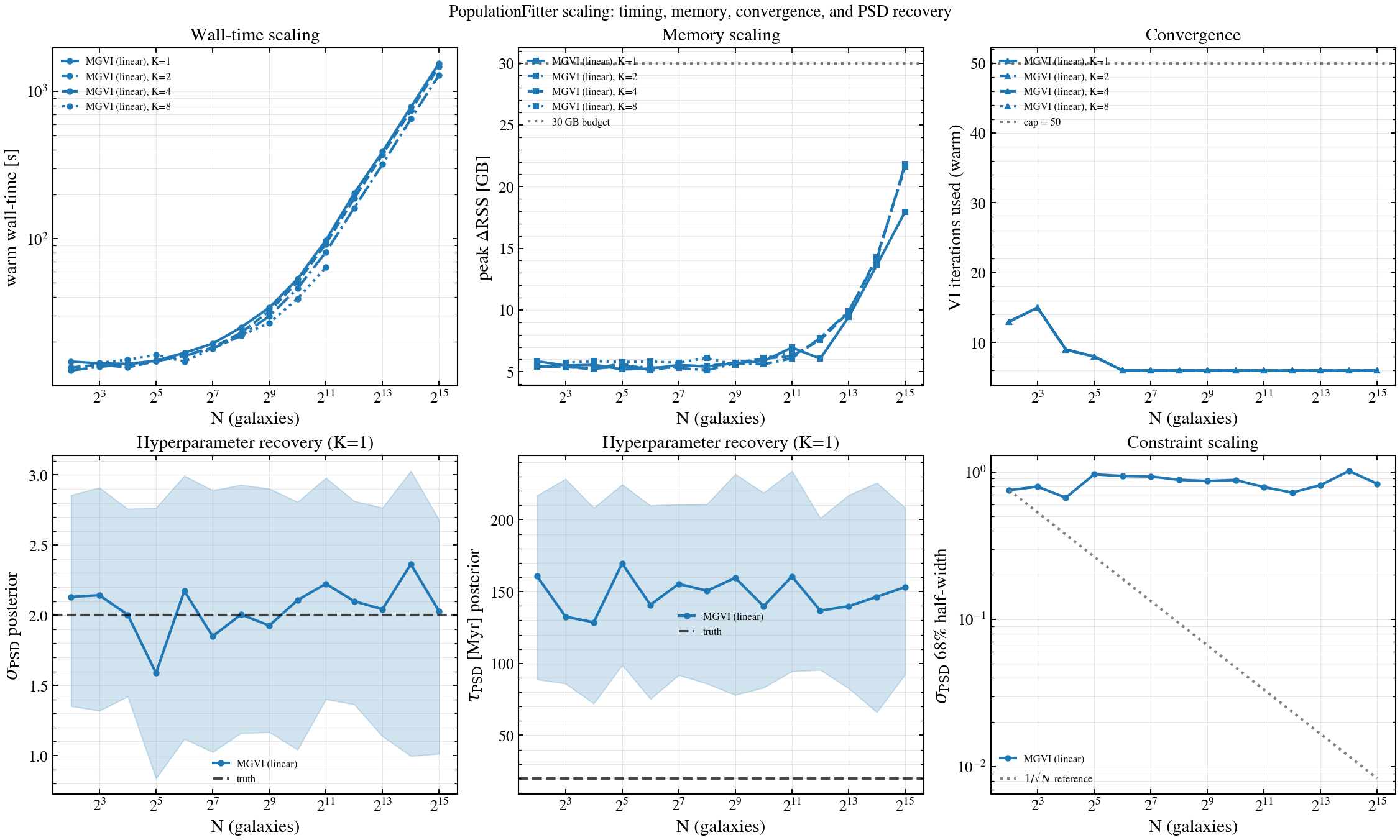
Renders the wall-time / peak-memory / iteration scaling of tengri’s two pure-JAX population variational engines on a 5-band SDSS photometry catalog with a stochastic-SFH forward model:
native_vi_linear— linearised geometric VI (MGVI).native_vi_nonlinear— full geometric VI (geoVI).
The grid scans N ∈ {4, …, 8192} galaxies × K ∈ {1, 2, 4, 8} forward chunks. Each cell is run in a fresh Python subprocess so the peak-RSS reading is clean.
This script is render-only: it loads results from
bench/results/vi_scaling_benchmark.json produced by
JAX_PLATFORMS=cpu python bench/scripts/benchmark_vi_xlarge.py
If the JSON is absent, the script prints instructions and exits.
Convergence policy¶
Both engines stop early via kl_rtol=1e-2. The benchmark uses an
iteration cap of 50 (retry 100); a row is converged iff
iters_used < cap. Non-converged rows are flagged on the iteration
panel.
Memory policy¶
Each run is bounded to ≤ 30 GB peak per worker. Columns abort once a row exceeds the budget — those (N, K) cells are absent from the plot.
from __future__ import annotations
import json
from pathlib import Path
import matplotlib.pyplot as plt
import numpy as np
from tengri import setup_style
setup_style()
def _find_results() -> Path | None:
"""Locate the benchmark JSON from project root or docs build cwd."""
name = "vi_scaling_benchmark.json"
for p in [
Path("data") / name,
Path("../data") / name,
Path("../../data") / name,
Path("../../../data") / name,
]:
if p.exists():
return p
return None
RESULTS = _find_results()
if RESULTS is None:
# Render-only example: never run the benchmark from a docs build.
# If the cached JSON is absent, emit a placeholder figure with
# instructions so the gallery still renders cleanly.
fig, ax = plt.subplots(figsize=(8, 4))
ax.axis("off")
ax.text(
0.5,
0.5,
"Population VI scaling benchmark not yet generated.\n\n"
"Produce the cached results once with:\n\n"
" JAX_PLATFORMS=cpu python bench/scripts/benchmark_vi_xlarge.py\n\n"
"It writes bench/results/vi_scaling_benchmark.json, which this gallery\n"
"script then renders without re-running the benchmark.",
ha="center",
va="center",
fontsize=11,
family="monospace",
bbox=dict(boxstyle="round,pad=0.7", fc="#f6f6f6", ec="#999"),
)
plt.show()
raise SystemExit(0)
with RESULTS.open() as f:
rows = json.load(f)
# Drop error rows and TIMEOUTs from the plot grid.
rows = [r for r in rows if not r.get("error") and r.get("wall_s_warm", -1) > 0]
methods = ("native_vi_linear", "native_vi_nonlinear")
labels = {"native_vi_linear": "MGVI (linear)", "native_vi_nonlinear": "geoVI (nonlinear)"}
colors = {"native_vi_linear": "#1f77b4", "native_vi_nonlinear": "#d62728"}
ks = sorted({r["forward_chunk_size"] for r in rows})
linestyles = {1: "-", 2: "--", 4: "-.", 8: ":"}
def _series(method: str, k: int, key: str) -> tuple[np.ndarray, np.ndarray]:
sel = [r for r in rows if r["method"] == method and r["forward_chunk_size"] == k]
sel.sort(key=lambda r: r["n_gal"])
if not sel:
return np.array([]), np.array([])
return (
np.array([r["n_gal"] for r in sel]),
np.array([r[key] for r in sel], dtype=float),
)
fig, axes = plt.subplots(2, 3, figsize=(15, 9), constrained_layout=True)
ax_t, ax_m, ax_i = axes[0]
ax_sig, ax_tau, ax_sig_err = axes[1]
# --- Panel 1: warm wall-time vs N ---
for method in methods:
for k in ks:
n, t = _series(method, k, "wall_s_warm")
if n.size == 0:
continue
ax_t.plot(
n,
t,
color=colors[method],
linestyle=linestyles.get(k, "-"),
marker="o",
markersize=4,
label=f"{labels[method]}, K={k}",
)
ax_t.set_xscale("log", base=2)
ax_t.set_yscale("log")
ax_t.set_xlabel("N (galaxies)")
ax_t.set_ylabel("warm wall-time [s]")
ax_t.set_title("Wall-time scaling")
ax_t.grid(True, which="both", alpha=0.3)
ax_t.legend(fontsize=8, loc="upper left")
# --- Panel 2: ΔRSS vs N ---
for method in methods:
for k in ks:
n, m = _series(method, k, "rss_delta_gb")
if n.size == 0:
continue
ax_m.plot(
n,
m,
color=colors[method],
linestyle=linestyles.get(k, "-"),
marker="s",
markersize=4,
label=f"{labels[method]}, K={k}",
)
ax_m.axhline(30.0, color="k", linestyle=":", alpha=0.5, label="30 GB budget")
ax_m.set_xscale("log", base=2)
ax_m.set_xlabel("N (galaxies)")
ax_m.set_ylabel("peak ΔRSS [GB]")
ax_m.set_title("Memory scaling")
ax_m.grid(True, which="both", alpha=0.3)
ax_m.legend(fontsize=8, loc="upper left")
# --- Panel 3: VI iterations to convergence ---
for method in methods:
for k in ks:
n, it = _series(method, k, "n_iters_used_warm")
if n.size == 0:
continue
ax_i.plot(
n,
it,
color=colors[method],
linestyle=linestyles.get(k, "-"),
marker="^",
markersize=4,
label=f"{labels[method]}, K={k}",
)
# Mark non-converged rows.
not_conv = [r for r in rows if not r.get("converged", False)]
if not_conv:
ax_i.scatter(
[r["n_gal"] for r in not_conv],
[r["n_iters_used_warm"] for r in not_conv],
marker="x",
color="red",
s=80,
zorder=5,
label="hit cap (NOT converged)",
)
cap = max((r["n_iters_max"] for r in rows), default=50)
ax_i.axhline(cap, color="k", linestyle=":", alpha=0.5, label=f"cap = {cap}")
ax_i.set_xscale("log", base=2)
ax_i.set_xlabel("N (galaxies)")
ax_i.set_ylabel("VI iterations used (warm)")
ax_i.set_title("Convergence")
ax_i.grid(True, which="both", alpha=0.3)
ax_i.legend(fontsize=8, loc="upper left")
# --- Panels 4 & 5: σ_PSD and τ_PSD constraint vs N (K=1 only) ---
TRUTH_SIGMA = 2.0
TRUTH_TAU = 20.0
def _constraint_series(
method: str, key: str
) -> tuple[np.ndarray, np.ndarray, np.ndarray, np.ndarray]:
sel = [
r for r in rows if r["method"] == method and r["forward_chunk_size"] == 1 and r.get(key)
]
sel.sort(key=lambda r: r["n_gal"])
if not sel:
return (np.array([]),) * 4
n = np.array([r["n_gal"] for r in sel])
med = np.array([r[key]["median"] for r in sel])
p16 = np.array([r[key]["p16"] for r in sel])
p84 = np.array([r[key]["p84"] for r in sel])
return n, med, p16, p84
for ax, (key, truth, ylabel) in zip(
(ax_sig, ax_tau),
(
("psd_sigma_summary", TRUTH_SIGMA, r"$\sigma_{\rm PSD}$ posterior"),
("psd_tau_summary", TRUTH_TAU, r"$\tau_{\rm PSD}$ [Myr] posterior"),
),
):
for method in methods:
n, med, p16, p84 = _constraint_series(method, key)
if n.size == 0:
continue
ax.fill_between(n, p16, p84, color=colors[method], alpha=0.2)
ax.plot(n, med, color=colors[method], marker="o", markersize=4, label=labels[method])
ax.axhline(truth, color="k", linestyle="--", alpha=0.7, label="truth")
ax.set_xscale("log", base=2)
ax.set_xlabel("N (galaxies)")
ax.set_ylabel(ylabel)
ax.set_title("Hyperparameter recovery (K=1)")
ax.grid(True, which="both", alpha=0.3)
ax.legend(fontsize=8, loc="best")
# --- Panel 6: σ posterior std vs N (Cramér–Rao-like 1/sqrt(N) reference) ---
for method in methods:
n, med, p16, p84 = _constraint_series(method, "psd_sigma_summary")
if n.size == 0:
continue
width = (p84 - p16) / 2.0 # ~1σ half-width
ax_sig_err.plot(
n,
width,
color=colors[method],
marker="o",
markersize=4,
label=f"{labels[method]}",
)
# 1/sqrt(N) reference, normalized to first MGVI point if available.
ref_n, _, p16r, p84r = _constraint_series("native_vi_linear", "psd_sigma_summary")
if ref_n.size > 0:
w0 = (p84r[0] - p16r[0]) / 2.0
ref = w0 * np.sqrt(ref_n[0] / ref_n)
ax_sig_err.plot(ref_n, ref, color="gray", linestyle=":", label=r"$1/\sqrt{N}$ reference")
ax_sig_err.set_xscale("log", base=2)
ax_sig_err.set_yscale("log")
ax_sig_err.set_xlabel("N (galaxies)")
ax_sig_err.set_ylabel(r"$\sigma_{\rm PSD}$ 68% half-width")
ax_sig_err.set_title("Constraint scaling")
ax_sig_err.grid(True, which="both", alpha=0.3)
ax_sig_err.legend(fontsize=8, loc="best")
fig.suptitle(
"PopulationFitter scaling: timing, memory, convergence, and PSD recovery",
fontsize=13,
)
plt.savefig("plot_population_scaling.png", dpi=150, bbox_inches="tight")
plt.show()