Note
Go to the end to download the full example code.
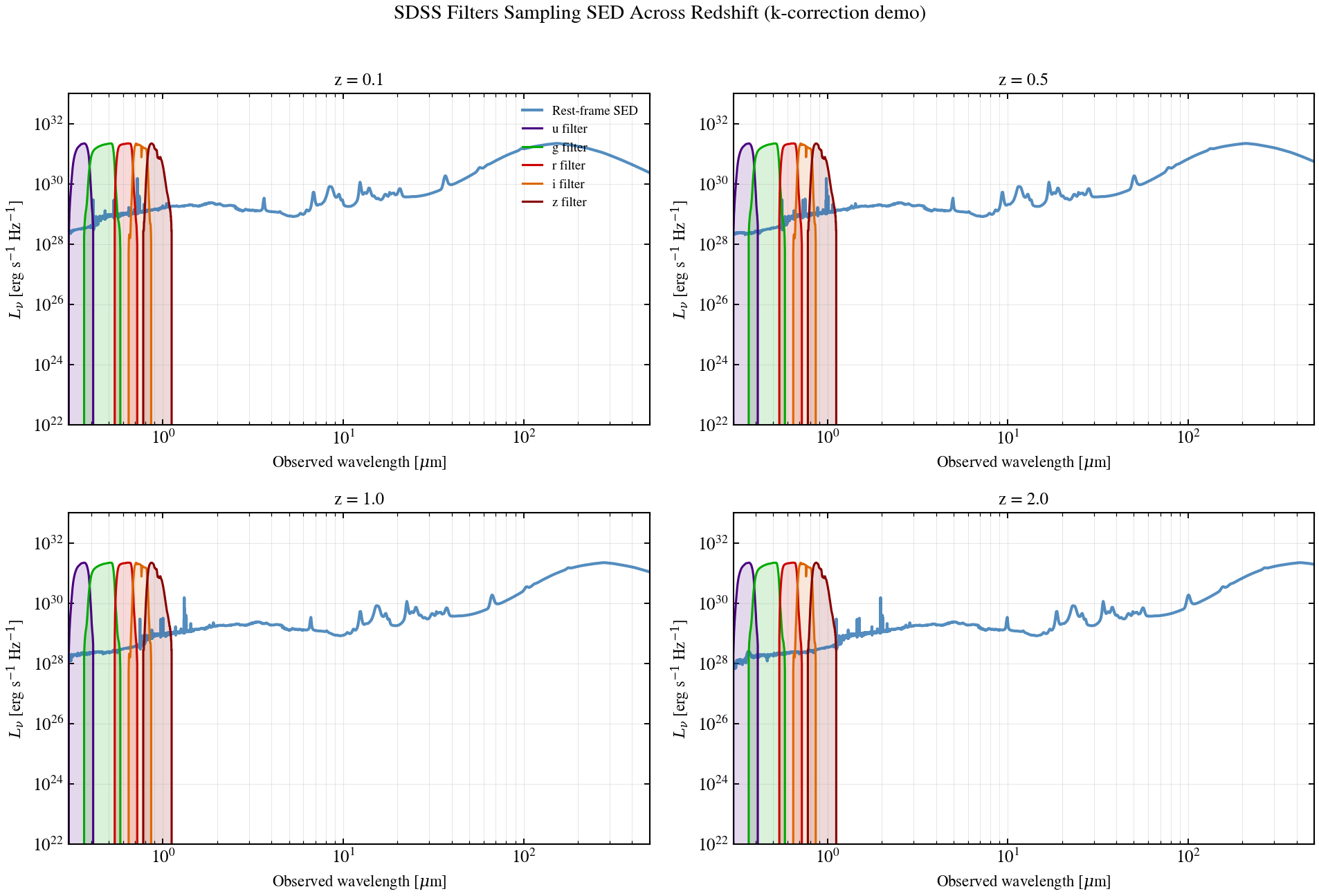
Filter Sampling Across Redshift¶
Rest-frame stellar continuum overlaid with redshifted SDSS ugriz transmission curves at z ∈ {0.1, 0.5, 1.0, 2.0}. The plot shows which features each band actually samples as a galaxy moves out — the textbook source of k-correction sign.
from pathlib import Path
import jax
import jax.numpy as jnp
import matplotlib
matplotlib.use("Agg")
import matplotlib.pyplot as plt
import numpy as np
jax.config.update("jax_enable_x64", True)
from tengri import (
Fixed,
Observation,
Parameters,
SEDModel,
Spectroscopy,
load_filter_set,
load_ssp_data,
setup_style,
)
setup_style()
def _find_ssp():
"""Find SSP data file in standard locations."""
name = "ssp_prsc_miles_chabrier_wNE_logGasU-3.0_logGasZ0.0.h5"
for p in [
Path("data") / name,
Path("../data") / name,
Path("../../data") / name,
Path("../../../data") / name,
]:
if p.exists():
return str(p)
return None
def _find_filters():
"""Find filter cache directory in standard locations."""
for p in [
Path("data/filters"),
Path("../data/filters"),
Path("../../data/filters"),
Path("../../../data/filters"),
]:
if p.exists():
return str(p)
return "data/filters"
ssp_path = _find_ssp()
if ssp_path is None:
raise FileNotFoundError("SSP data not found — skipping example")
filter_dir = _find_filters()
ssp = load_ssp_data(ssp_path)
# Generate a template star-forming galaxy SED (rest-frame)
wave_rest = jnp.logspace(jnp.log10(1000.0), jnp.log10(3e5), 500) # 0.1 µm – 30 µm [Å]
obs_dummy = Observation(spectroscopy=Spectroscopy(wave_obs=wave_rest))
spec = Parameters(
mean_sfh_type="tsnorm",
dust_emission="draine_li2007",
sfh_tsnorm_log_peak_sfr=Fixed(1.0),
sfh_tsnorm_peak_lbt_gyr=Fixed(2.0),
sfh_tsnorm_width_gyr=Fixed(1.5),
sfh_tsnorm_skew=Fixed(0.0),
sfh_tsnorm_trunc=Fixed(2.0),
met_logzsol=Fixed(0.0),
dust_tau_bc=Fixed(0.3),
dust_tau_diff=Fixed(0.2),
dust_slope=Fixed(-0.7),
dust_umin=Fixed(2.0),
dust_qpah=Fixed(3.5),
dust_gamma_dl=Fixed(0.02),
redshift=Fixed(0.0), # Rest-frame for now
)
model = SEDModel(spec, ssp, observation=obs_dummy)
key = jax.random.PRNGKey(42)
params = spec.sample(key)
pred = model.predict_rest_sed(params)
wave_rest_um = np.array(pred.wavelength) / 1e4
sed_rest = np.array(pred.sed)
# Load SDSS filters
filter_names = ["sdss_u", "sdss_g", "sdss_r", "sdss_i", "sdss_z"]
_, _, filter_curves = load_filter_set(filter_names, cache_dir=filter_dir)
band_colors = {
"sdss_u": "#4B0082",
"sdss_g": "#00AA00",
"sdss_r": "#CC0000",
"sdss_i": "#DD6600",
"sdss_z": "#880000",
}
# Redshifts to visualize
redshifts = [0.1, 0.5, 1.0, 2.0]
fig, axes = plt.subplots(2, 2, figsize=(13, 9))
axes = axes.flatten()
for i, z in enumerate(redshifts):
ax = axes[i]
# Plot rest-frame SED (shifted to observed frame)
wave_obs_um = wave_rest_um * (1 + z)
ax.loglog(
wave_obs_um,
sed_rest,
color="C0",
lw=2.0,
label="Rest-frame SED",
alpha=0.7,
)
# Overlay redshifted filters
for fc, fname in zip(filter_curves, filter_names):
wave_filter = np.array(fc.wave) / 1e4 # Å → µm
trans = np.array(fc.trans)
# Normalize transmission to fit on log plot
trans_scaled = trans * np.max(sed_rest) / np.max(trans)
color = band_colors[fname]
band_short = fname.replace("sdss_", "")
ax.fill_between(wave_filter, 1e20, trans_scaled, alpha=0.15, color=color)
ax.plot(wave_filter, trans_scaled, color=color, lw=1.5, label=f"{band_short} filter")
ax.set_xlim(0.3, 5e2)
ax.set_ylim(1e22, 1e33)
ax.set_xlabel(r"Observed wavelength [$\mu$m]", fontsize=11)
ax.set_ylabel(r"$L_\nu$ [erg s$^{-1}$ Hz$^{-1}$]", fontsize=11)
ax.set_title(f"z = {z}", fontsize=12)
if i == 0:
ax.legend(fontsize=9, frameon=False, loc="upper right")
ax.grid(True, alpha=0.3, which="both")
fig.suptitle("SDSS Filters Sampling SED Across Redshift (k-correction demo)", fontsize=14)
fig.tight_layout(rect=[0, 0.01, 1, 0.97])
# Save to script directory
script_dir = Path(__file__).resolve().parent if "__file__" in dir() else Path(".")
plt.savefig(str(script_dir / "plot_redshift_filter_grid.png"), dpi=150, bbox_inches="tight")
plt.close()