Note
Go to the end to download the full example code.
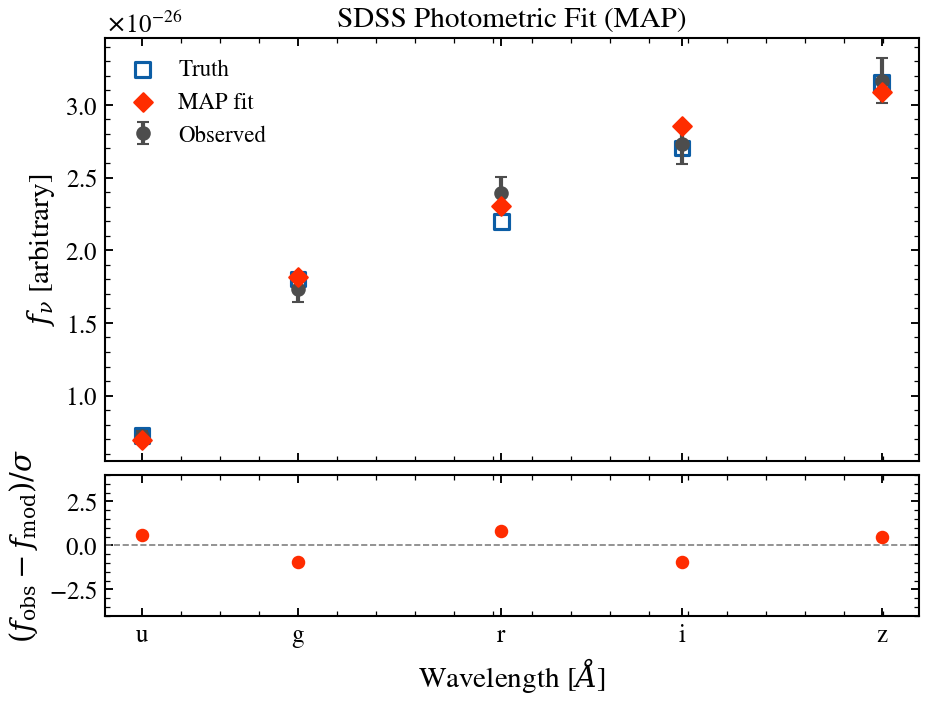
Photometric SED Fit¶
Generate a mock galaxy with SDSS ugriz photometry and fit it using tengri’s variational inference. Shows observed vs model photometry with error bars and residuals.
from pathlib import Path
import jax
import matplotlib.pyplot as plt
import numpy as np
from tengri import (
Fitter,
Fixed,
Observation,
Parameters,
Photometry,
SEDModel,
Uniform,
load_ssp_data,
)
from tengri.analysis.plotting import setup_style
setup_style()
# --- Check for SSP data ---
def _find_ssp():
"""Locate SSP data from project root or docs/ (sphinx-gallery) cwd."""
name = "ssp_prsc_miles_chabrier_wNE_logGasU-3.0_logGasZ0.0.h5"
for p in [
Path("data") / name,
Path("../data") / name,
Path("../../data") / name,
Path("../../../data") / name,
]:
if p.exists():
return str(p)
return None
SSP_PATH = _find_ssp()
# Locate filter cache
_FILTER_DIR = next(
(
str(d)
for d in [
Path("data/filters"),
Path("../data/filters"),
Path("../../data/filters"),
Path("../../../data/filters"),
]
if d.exists()
),
"data/filters",
)
if SSP_PATH is None:
raise FileNotFoundError("SSP data not found — skipping example")
# --- Setup ---
ssp_data = load_ssp_data(SSP_PATH)
bands = ["sdss_u", "sdss_g", "sdss_r", "sdss_i", "sdss_z"]
obs = Observation(
photometry=Photometry.from_names(bands, cache_dir=_FILTER_DIR),
)
spec = Parameters(
sfh_tsnorm_log_peak_sfr=Uniform(-1.0, 2.5),
sfh_tsnorm_peak_lbt_gyr=Uniform(0.5, 12.0),
sfh_tsnorm_width_gyr=Uniform(0.3, 5.0),
sfh_tsnorm_skew=Uniform(-3.0, 3.0),
sfh_tsnorm_trunc=Uniform(1.0, 10.0),
met_logzsol=Uniform(-2.0, 0.2),
dust_tau_bc=Fixed(0.3),
dust_tau_diff=Fixed(0.2),
dust_slope=Fixed(-0.7),
redshift=Fixed(0.05),
)
model = SEDModel(spec, ssp_data, observation=obs)
# --- Generate mock data (star-forming galaxy) ---
true_params = spec.sample(jax.random.PRNGKey(42))
true_params["sfh_tsnorm_peak_lbt_gyr"] = 3.0
true_params["sfh_tsnorm_width_gyr"] = 2.0
true_params["sfh_tsnorm_log_peak_sfr"] = 1.0
true_params["sfh_tsnorm_skew"] = 0.3 # Positive skew = recent star formation
mock = model.mock(true_params, snr=20.0, key=jax.random.PRNGKey(0))
# --- Fit with MAP ---
fitter = Fitter(model, mock.flux_obs, mock.noise)
posterior = fitter.run("map", optimizer="adam", n_steps=300, verbose=False)
best_fit = model.predict_photometry(posterior.params)
# --- Plot ---
wave_eff = np.array([3551, 4686, 6166, 7480, 8932]) # SDSS effective wavelengths
band_names = ["u", "g", "r", "i", "z"]
fig, (ax, ax_res) = plt.subplots(
2, 1, figsize=(7, 5), height_ratios=[3, 1], sharex=True, gridspec_kw={"hspace": 0.05}
)
ax.errorbar(
wave_eff,
np.array(mock.flux_obs),
yerr=np.array(mock.noise),
fmt="o",
color="0.3",
ms=6,
capsize=3,
label="Observed",
zorder=5,
)
ax.scatter(
wave_eff,
np.array(mock.flux_true),
marker="s",
s=50,
facecolors="none",
edgecolors="C0",
lw=1.5,
label="Truth",
zorder=4,
)
ax.scatter(wave_eff, np.array(best_fit), marker="D", s=40, color="C3", label="MAP fit", zorder=6)
ax.set_ylabel(r"$f_\nu$ [arbitrary]")
ax.legend(frameon=False)
ax.set_title("SDSS Photometric Fit (MAP)")
residuals = (np.array(mock.flux_obs) - np.array(best_fit)) / np.array(mock.noise)
ax_res.axhline(0, color="0.5", ls="--", lw=0.8)
ax_res.scatter(wave_eff, residuals, c="C3", s=30, zorder=5)
ax_res.set_xlabel(r"Wavelength [$\AA$]")
ax_res.set_ylabel(r"$(f_\mathrm{obs} - f_\mathrm{mod}) / \sigma$")
ax_res.set_ylim(-4, 4)
ax_res.set_xticks(wave_eff)
ax_res.set_xticklabels(band_names)
fig.tight_layout()
plt.savefig("plot_photometric_fit.png", dpi=150, bbox_inches="tight")
plt.show()