Note
Go to the end to download the full example code.
Hierarchical PSD Inference¶
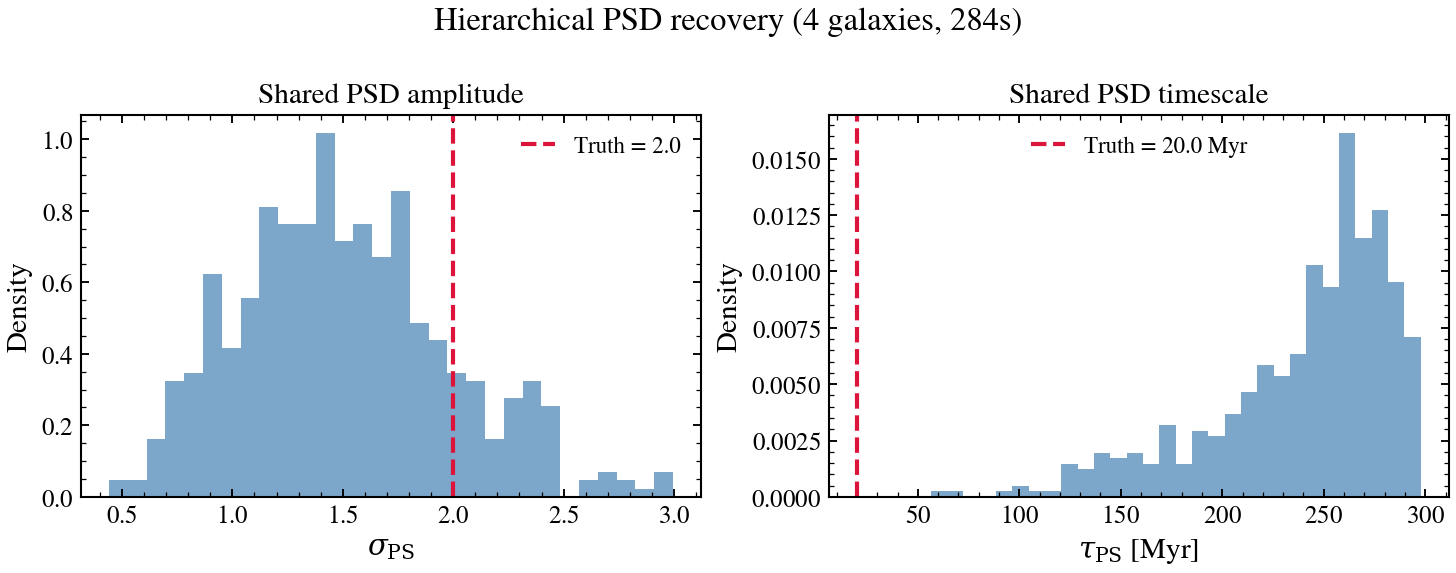
Sets up a small population of mock galaxies sharing the same burstiness PSD parameters (sigma, tau), runs PopulationFitter briefly, and displays the shared PSD posterior vs truth.
import time
from pathlib import Path
import jax
import matplotlib.pyplot as plt
import numpy as np
jax.config.update("jax_enable_x64", True)
from tengri import (
Fixed,
Observation,
Parameters,
Photometry,
PopulationFitter,
SEDModel,
Uniform,
load_ssp_data,
setup_style,
)
setup_style()
# --- Data ---
def _find_ssp():
"""Locate SSP data from project root or docs/ (sphinx-gallery) cwd."""
name = "ssp_prsc_miles_chabrier_wNE_logGasU-3.0_logGasZ0.0.h5"
for p in [
Path("data") / name,
Path("../data") / name,
Path("../../data") / name,
Path("../../../data") / name,
]:
if p.exists():
return str(p)
return None
SSP_PATH = _find_ssp()
if SSP_PATH is None:
raise FileNotFoundError("SSP data not found — skipping example")
ssp = load_ssp_data(SSP_PATH)
obs = Observation(
photometry=Photometry.from_names(["sdss_u", "sdss_g", "sdss_r", "sdss_i", "sdss_z"])
)
# --- True shared PSD ---
TRUE_SIGMA = 2.0
TRUE_TAU = 20.0
N_GAL = 4
def model_factory(psd_sigma=1.0, psd_tau_myr=50.0):
"""Create a SEDModel with fixed PSD — called by PopulationFitter."""
spec = Parameters(
sfh_tsnorm_log_peak_sfr=Uniform(-1.0, 2.5),
sfh_tsnorm_peak_lbt_gyr=Uniform(0.5, 12.0),
sfh_tsnorm_width_gyr=Uniform(0.3, 5.0),
sfh_tsnorm_skew=Uniform(-3.0, 3.0),
sfh_tsnorm_trunc=Uniform(1.0, 10.0),
sfh_field_psd_sigma=Fixed(psd_sigma),
sfh_field_psd_tau_myr=Fixed(psd_tau_myr),
met_logzsol=Uniform(-2.0, 0.2),
dust_tau_bc=Uniform(0.0, 2.0),
dust_tau_diff=Uniform(0.0, 1.5),
dust_slope=Fixed(-0.7),
redshift=Fixed(0.1),
mean_sfh_type=["tsnorm", "field"],
n_grid=128,
)
return SEDModel(spec, ssp, observation=obs)
# --- Generate mock galaxies ---
key = jax.random.PRNGKey(42)
model_gen = model_factory(psd_sigma=TRUE_SIGMA, psd_tau_myr=TRUE_TAU)
galaxies = []
for i in range(N_GAL):
k = jax.random.fold_in(key, i)
params = model_gen.spec.sample(k)
mock = model_gen.mock(params, snr=20.0, key=jax.random.fold_in(k, 1))
galaxies.append({"flux_obs": mock.flux_obs, "noise": mock.noise})
print(f"Generated {N_GAL} mock galaxies with sigma={TRUE_SIGMA}, tau={TRUE_TAU} Myr")
# --- Hierarchical fit (quick) ---
hfitter = PopulationFitter(
model_factory,
galaxies,
psd_sigma_prior=(0.1, 4.0),
psd_tau_prior=(1.0, 300.0),
)
t0 = time.perf_counter()
result = hfitter.run(
"vi_linear",
n_iterations=20,
n_samples=4,
n_posterior_samples=500,
verbose=False,
key=jax.random.PRNGKey(0),
)
elapsed = time.perf_counter() - t0
print(f"Hierarchical fit: {elapsed:.1f}s")
# --- Figure: shared PSD posterior ---
# shared_samples keys depend on the spec — print to be safe and pick the
# psd amplitude/timescale entries by substring match.
keys = list(result.shared_samples.keys())
sig_key = next(k for k in keys if "psd" in k and ("sigma" in k or "_u" in k or "amp" in k))
tau_key = next(k for k in keys if "psd" in k and ("tau" in k))
sig_samples = np.array(result.shared_samples[sig_key])
tau_samples = np.array(result.shared_samples[tau_key])
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(10, 4))
ax1.hist(sig_samples, bins=30, density=True, alpha=0.7, color="steelblue")
ax1.axvline(TRUE_SIGMA, color="crimson", ls="--", lw=2, label=f"Truth = {TRUE_SIGMA}")
ax1.set_xlabel(r"$\sigma_{\rm PS}$")
ax1.set_ylabel("Density")
ax1.set_title("Shared PSD amplitude")
ax1.legend()
ax2.hist(tau_samples, bins=30, density=True, alpha=0.7, color="steelblue")
ax2.axvline(TRUE_TAU, color="crimson", ls="--", lw=2, label=f"Truth = {TRUE_TAU} Myr")
ax2.set_xlabel(r"$\tau_{\rm PS}$ [Myr]")
ax2.set_ylabel("Density")
ax2.set_title("Shared PSD timescale")
ax2.legend()
fig.suptitle(f"Hierarchical PSD recovery ({N_GAL} galaxies, {elapsed:.0f}s)")
fig.tight_layout()
outdir = Path(__file__).resolve().parent.parent / "figures" if "__file__" in dir() else Path(".")
outdir.mkdir(parents=True, exist_ok=True)
plt.savefig("plot_hierarchical.png", dpi=150, bbox_inches="tight")
plt.show()