Note
Go to the end to download the full example code.
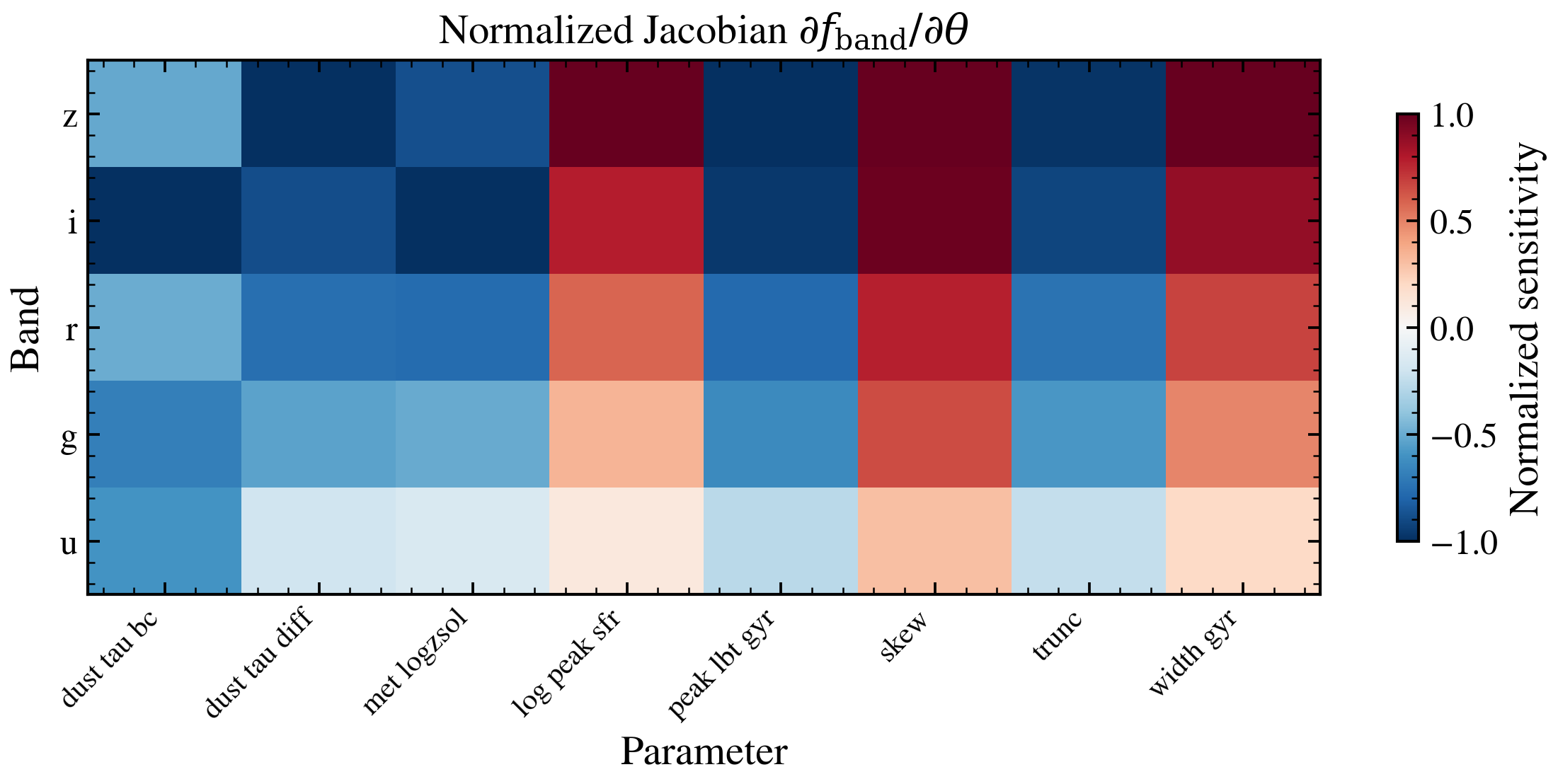
Gradient Sensitivity Heatmap¶
Computes the Jacobian d(flux)/d(theta) of the forward model and displays it as a heatmap showing which photometric bands are sensitive to which physical parameters.
from pathlib import Path
import jax
import jax.numpy as jnp
import matplotlib.pyplot as plt
import numpy as np
jax.config.update("jax_enable_x64", True)
from tengri import (
Fixed,
Observation,
Parameters,
Photometry,
SEDModel,
Uniform,
load_ssp_data,
setup_style,
)
setup_style()
# --- Data ---
def _find_ssp():
"""Locate SSP data from project root or docs/ (sphinx-gallery) cwd."""
name = "ssp_prsc_miles_chabrier_wNE_logGasU-3.0_logGasZ0.0.h5"
for p in [
Path("data") / name,
Path("../data") / name,
Path("../../data") / name,
Path("../../../data") / name,
]:
if p.exists():
return str(p)
return None
SSP_PATH = _find_ssp()
if SSP_PATH is None:
raise FileNotFoundError("SSP data not found — skipping example")
ssp = load_ssp_data(SSP_PATH)
bands = ["sdss_u", "sdss_g", "sdss_r", "sdss_i", "sdss_z"]
obs = Observation(photometry=Photometry.from_names(bands))
# --- SEDModel ---
spec = Parameters(
sfh_tsnorm_log_peak_sfr=Uniform(-1.0, 2.5),
sfh_tsnorm_peak_lbt_gyr=Uniform(0.5, 12.0),
sfh_tsnorm_width_gyr=Uniform(0.3, 5.0),
sfh_tsnorm_skew=Uniform(-3.0, 3.0),
sfh_tsnorm_trunc=Uniform(1.0, 10.0),
met_logzsol=Uniform(-2.0, 0.2),
dust_tau_bc=Uniform(0.0, 2.0),
dust_tau_diff=Uniform(0.0, 1.5),
dust_slope=Fixed(-0.7),
redshift=Fixed(0.1),
mean_sfh_type="tsnorm",
)
model = SEDModel(spec, ssp, observation=obs)
# --- Fiducial point ---
fiducial = {
"sfh_tsnorm_log_peak_sfr": 1.0,
"sfh_tsnorm_peak_lbt_gyr": 4.0,
"sfh_tsnorm_width_gyr": 2.0,
"sfh_tsnorm_skew": 0.0,
"sfh_tsnorm_trunc": 5.0,
"met_logzsol": -0.3,
"dust_tau_bc": 0.3,
"dust_tau_diff": 0.5,
"dust_slope": -0.7,
"redshift": 0.1,
}
free_names = spec.free_params
fixed_vals = spec.get_fixed_values()
def photometry_from_array(param_array):
"""Map flat array to photometric fluxes."""
params = dict(fixed_vals)
for i, name in enumerate(free_names):
params[name] = param_array[i]
return model.predict_photometry(params)
param_array = jnp.array([fiducial[n] for n in free_names])
jacobian = jax.jacobian(photometry_from_array)(param_array) # (n_bands, n_params)
# --- Figure: heatmap ---
J = np.array(jacobian)
# Normalize each column (parameter) to unit max for visual clarity
J_norm = J / (np.abs(J).max(axis=0, keepdims=True) + 1e-30)
fig, ax = plt.subplots(figsize=(8, 4))
short_names = [n.replace("sfh_tsnorm_", "").replace("_", " ") for n in free_names]
im = ax.imshow(J_norm, aspect="auto", cmap="RdBu_r", vmin=-1, vmax=1)
ax.set_xticks(range(len(free_names)))
ax.set_xticklabels(short_names, rotation=45, ha="right", fontsize=10)
ax.set_yticks(range(len(bands)))
ax.set_yticklabels([b.replace("sdss_", "") for b in bands])
ax.set_ylim(-0.5, len(bands) - 0.5)
ax.set_xlabel("Parameter")
ax.set_ylabel("Band")
ax.set_title(r"Normalized Jacobian $\partial f_{\rm band} / \partial \theta$")
fig.colorbar(im, ax=ax, shrink=0.8, label="Normalized sensitivity")
fig.tight_layout()
plt.savefig(
"plot_gradient_sensitivity.png",
dpi=150,
bbox_inches="tight",
)
plt.show()